
From de Bruijn Graphs to Rectangle Graphs for Genome Assembly
We came across an abstract that appears interesting, but we do not have access to the full paper. If you have a copy, would you please email it to us [samanta at homolog.us]?
Jigsaw puzzles were originally constructed by painting a picture on a rectangular piece of wood and further cutting it into smaller pieces with a jigsaw. The Jigsaw Puzzle Problem is to find an arrangement of these pieces that fills up the rectangle in such a way that neighboring pieces have matching boundaries with respect to color and texture. While the general Jigsaw Puzzle Problem is NP-complete [6], we discuss its simpler version (called Rectangle Puzzle Problem) and study the rectangle graphs, recently introduced by Bankevich et al., 2012 [3], for assembling such puzzles. We establish the connection between Rectangle Puzzle Problem and the problem of assembling genomes from read-pairs, and further extend the analysis in [3] to real challenges encountered in applications of rectangle graphs in genome assembly. We demonstrate that addressing these challenges results in an assembler SPAdes+ that improves on existing assembly algorithms in the case of bacterial genomes (including particularly difficult case of genome assemblies from single cells).
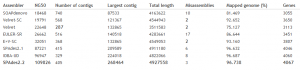
The approach is integrated into SPAdes Genome Assembler developed by Algorithmic Biology Lab, St. Petersburg, Russia. Their posted benchmarks on E. coli assemble appear impressive.
If nobody mails us the paper, we threaten to start reading the SPades source- code (and kill us in the process :) ).