The 'Right Way' and the 'Wrong Way' to Criticize Science
-————————————————————–
Lior Pachter
Every time we go to Lior Pachter’s blog, we start with the assumption that he is making a highly inflated claim. His titles seem too aggressive and the accusations appear too strong. Moreover, his posts are loaded with information and require more than one reading, whereas the time-saving approach is to check what other scientists are saying in twitter. There you find nine out of ten disagree with him and so he must be wrong !!
If, instead, you take enough care to go through all his points, it becomes quite clear that he is factually correct and is making claims justified by enough evidence. Take the current example, where he accuses Manolis Kellis of MIT of fraud and misconduct. Those are very strong words in academia and we started reading Pachter’s blog with the default assumption that Kellis made an honest mistake. Unfortunately, after we checked all details provided in Pachter’s post, it became clear that Kellis engaged in unethical and deceptive practices well outside the norms of standard academic procedure.
What did he do? In a nutshell, he surreptitiously updated parts of supplement of a paper published in Nature Biotech months after publication, but did not declare that change anywhere. The updated supplement came with the following innocuous note.
In the version of this file originally posted online, in equation (12) in Supplementary Note 1, the word max should have been min. The correct formula was implemented in the source code. Clarification has been made to Supplementary Notes 1.1, 1.3, 1.6 and Supplementary Figure 4 about
the practical implementation of network deconvolution and parameter selection for application to the examples used in the paper. In Supplementary Data, in the file ND.m on line 55, the parameter delta should have been set to 1 epsilon (that is, delta = 1 0.01), rather than 1. These errors have been corrected in this file and in the Supplementary Data zip file as of 26 August 2013.
So far so good, but if one compares the old supplement with the new one, one discovers more significant changes being made than what was stated in the note. For example, Fig. S4 changed qualitatively and its caption changed from -
As it is illustrated in this ?gure, choosing ? close to one (i.e., considering higher order indirect interactions) leads to best performance in all considered network deconvolution applications. Further, note that despite the social network application, considering higher order indirect interactions is important in gene regulatory inference and protein structural constraint inference applications.
to -
For regulatory network inference, we used ? = 0.5, for protein contact maps, we used ? = 0.99, and for co-authorship network, we used ? = 0.95. Further, note that despite the social network application, considering higher order indirect interactions is important in gene regulatory inference and protein structural constraint inference applications.
That completely changes the implication of the figure, and Lior Pachter discussed mathematical reason behind the above modification. Irrespective of whether his math is right, there are two even more fundamental problems in surreptitiously switching figures.
a) Clearly, with the above changes, the updated version of the paper is qualitatively different from the version that went through peer review. In other words, one can easily argue that the currently published paper is NOT peer-reviewed. If everyone is allowed to present one narrative during the review process and another one months after publication, the scientific publication process will turn into a farce.
b) In the past, occasionally scientists were allowed to correct or modify erroneous figures after publication. The usual procedure is to tell the editor and issue an erratum. The erratum includes a note saying that the corrected figure does not change the science presented in the paper, and if the editor has any doubt, he consults the reviewers about the claim. Instead Kellis merely sneaked in the new supplementary figure without mentioning about it anywhere. We are not sure, why such practices should be condoned.
-—————————————————————-
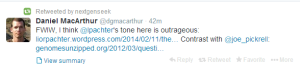
Joe Pickrell
After reading Joe Pickrell’s commentary about recent Elhaik/Graur paper, we came to conclude that he is seriously compromised as a scientist.
Last month, Eran Elhaik, Dan Graur and co-authors published a paper, where they suggested that a leading researcher in their sub-field of human population genetics used statistics to lie. Based on their analysis, the paper by Mendez et al. inflated a bunch of numbers to make a newsworthy discovery. Those could be honest mistakes and we specifically asked the first author Eran Elhaik whether that is the case.
Q6. Based on your analysis, do the statistical errors appear to be intentional (purposely done to make inflated claim), or are they likely to be honest mistakes?
EE: I cannot speak to intent, however random errors are likely to both inflate and deflate the chromosome age and this is not what we observed.
For a second opinion, we headed to Joe Pickrell’s blog, where he claimed to have discussed Graur/Mendez paper. Sadly, we were very disappointed. He seemed to have completely avoided the main point of Graur/Elhaik paper (accusation of purposely pushing numbers to make inflated claims), which appears to be very serious to us for integrity of the entire field. It was even more strange, that he completely avoided the question after repeated prodding by us and Dan Graur.
Joe Pickrell is an expert in this area and he also maintains Haldane’s Sieve blog, whose presumed role is to provide an open online discussion forum for preprints in population genetics. How can an expert in a field not care about deceptive behavior bordering on fraud? What kind of science does he stand for?
Interestingly, Joe Pickrell does not see anything wrong in switching figures post-publication either.
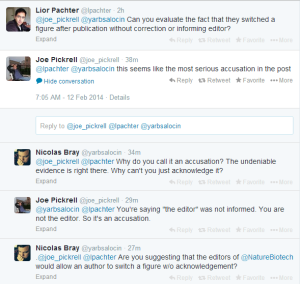
-————————————————–
Edit. 2/19
In a twitter conversation, Joe Pickrell provided his objective criticism of Elhaik vs Mendez paper. For the sake of completeness and readers’ benefit, we add them here. Thank you Joe !!