
Introducing 'Pandora's Toolbox' and 'Pandora's Modules'
Pandora’s Toolbox
Github link - here
License - GNU GPL3
Description -
Pandora’s Toolbox is a collection of source codes of well-known bioinformatics programs, all at the same place. You can now run ‘git clone’ and then ‘make’ once at the main folder. That will compile everything and collect executables in the ‘bin’ folder.
Previously, every time we tried to build them, we faced the problem of having to collect the source codes from various websites. Then each program had to be compiled individually, and some of those using boost were nightmares to compile. Also, many programs have interdependencies and we ended up having five copies of different versions of BWA or samtools codes.
Our collection reduces those dependencies and makes life easier for those working on different bioinformatics programs. The collection includes a boost_1_55_0 folder to remove external boost dependencies. We are working on sorting out all bwa and other inter-dependencies.
The following programs are included in the current version. Please DO NOT cite us, but cite the authors of individual programs.
1. KMC - kmer counter
Cite Deorowitz et al..
2. bwa - aligner
Cite Heng li.
3. DALIGNER - pacbio aligner
Cite Gene Myers.
4. HMMER3
Cite Sean Eddy’s PLOS paper -
http://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.100219 5
We cannot call it HMMER, because Sean Eddy and Janelia lawyers made such
a thing illegal by copyrighting trademarking the HMMER name.
5. bcalm - de Bruijn graph compressor
Cite Chikhi et al.
6. Minia - assembler
Cite Chikhi et al.
7. samtools - NGS data handling
Cite Heng Li.
8. SOAPdenovo2 - genome assembler
Cite Luo et al.
9. SOAPdenovo-trans - transcriptome assembler
Cite Xie et al.
10. SPAdes - genome assembler
Cite the SPAdes team.
11. Trinity - transcriptome assembler
Cite Grabher et al.
12. sailfish - RNAseq expression analysis
Cite Rob Patro et al..
[We are still working on compilation of this code.]
-————————————————————-
Pandora’s Modules
Github link - here
License - MIT
Description -
‘Pandora’s Modules’ includes various efficient code-blocks to be used by bioinformatics developers. Heng Li’s klib is a good example, minimizer being another. The only condition is that the library has to be accessible under MIT /BSD-style license, and therefore we cannot use a number of GPL3-licensed algorithms and have to re-code them. We are in very early stage of development with this collection, and will keep you posted as we incorporate various codes in it.
-————————————————————-
About the name
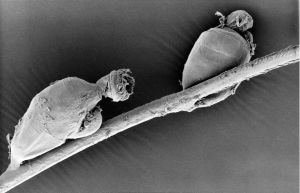
We named the modules after Symbion pandora, an unusual animal discovered by Danish researchers Peter Funch and R. M. Christensen in 1995. Many thanks to professor Funch for sharing the SEM image at the top of this post.
More on Symbion pandora -
Symbion pandora is a jug-shaped microscopic aquatic animal that dwells on the mouth-parts of Norway lobsters. The animals are less than mm wide, with sac-like bodies, and three distinctly different forms in different parts of their three-stage life cycle.
They are so unlike any known animal that its discovery by Danish scientists in 1995[1] led to the creation of a new phylum. The phylum Cycliophora, from the Greek for ‘carrying a small wheel’, was named after the creature’s circular mouth.
From their original Nature paper -
THE mouthparts of the Norway lobster Nephrops are colonized by an acoelomate metazoan, Symbion pandora gen. et sp. nov. Sessile stages continually produce inner buds replacing feeding structures. They also produce one of three motile stages: (1) larvae containing new feeding stages, (2) dwarf males, which settle on feeding stages, or (3) females, which settle onto lobster mouthparts, and eventually degenerate, giving rise to dispersive larvae. All motile stages are short-lived, and do not feed. The structure and function of the cilia suggest a phylogenetic position in Protostomia, while some aspects of inner budding and brooding of larvae are similar to those of Entoprocta and Ectoprocta. The dispersive larva possesses a mesodermal supporting chordoid structure, otherwise absent in protostomian larvae. We believe that all the above features of this previously undescribed species warrant the recognition of a new phylum with affinities to Ectoprocta and Entoprocta.