
Battle over #GWAS: Ken Weiss Edition
The battle over #GWAS continues, thus providing us with more opportunity to learn. Today’s finding is one of the best. Thanks to @dgmacarthur, we discovered the blog of professor Ken Weiss !
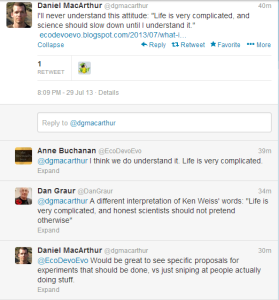
The blog post Daniel Macarthur referred to in above tweet can be accessed here. Kenneth M. Weiss is a professor of biology at Penn State University. In the above twitter battle, older people are deferential to him because of his contribution to science, and younger people are upset about his complaining. We decided to do neither, and chose to read his papers.
That is how we struck gold. This gentleman is wise beyond his years !!

He has been warning about false promises made by those pushing for large funding for GWAS since 2000, or possibly even before. Please take a look at his 2000 Nature commentary titled -
How many diseases does it take to map a gene with SNPs?
We are extracting few paragraphs below, but do encourage readers to go through the entire paper, because it is full of wisdom.
We do not contest that genome scanning for LD can work when there is a strong difference in the frequency of some risk genotype(s) between cases and controls. Nikali et al.70were able to map the gene responsible for a rare recessive ataxia in a Finnish cohort using an LD-based genome scan with microsatellite markers using only four affected individuals, demonstrating that it can, of course, work. Similarly, the poster boy of the SNP proponents wears a T-shirt advertising APOE ?4 and Alzheimer disease; this association is cited as proof-of-principle for SNP-based association mapping. By using a data set in which APOE ?4 was known to be over-represented in cases compared with controls, it has been possible to take a set of SNPs that are in LD with the risk allele and identify the association without studying APOE ?4 itself71,72. This case, stacked in favour of positive ?ndings, shows that neither coding SNPs, nor most SNPs, nor common SNPs, nor nearby SNPs, nor single SNPs, nor a SNP in a neighbouring gene, nor even simple haplotypes can be relied upon, a priori, as a single, casually designed source of LD markers. What this does show is the not-surprising fact that when there is an association with a single risk allele, one can identify this by using multiple markers which are in LD with this single risk allele14. How easily would genes such as BRCA1, BRCA2, PSEN1, PSEN2 and ABCA4, which have many alleles each of which increases risk of some common disease, be mapped by scrutinizing a set of common SNPs in a sample from a large cosmopolitan population13,14? No one can deny that disease pathways have been identi?ed by genetic epidemiology studies, making contributions to our biological knowledge. So far, however, it is lifestyle changes that have made the most impact on reduced (or increased!) incidence of chronic disease52,53,73. In those instances in which common variants affecting common disease really do exist, it is important to ?nd them. But we regularly hear grander promises, and it is at least fair to ask whether scaling up current genetic approaches, which have been likened to a search for a needle in a needle stack (a great many individually modest, effects), would be the wisest investment when a major justi?cation is that nothing else has worked so far.
To some this is an inapt question. New genes may identify previously unknown biochemical pathways that may lead to therapeutic pharmaceutical improvements (even if the latter are not speci?cally genetic). Other investigators argue that the identi?cation of even weakly predictive screening tools is suf?cient justi?cation, or hope that a worthwhile fraction of complex diseases will have common variants with individually modest effect but substantial attributable risk13,7. The level of merit in these arguments is dif?cult to evaluate, and the track record is unclear at best; but the arguments undeniably have aspects of rationalizing results that are not living up to expectations, sometimes based on unrealistically optimistic models20,21,74, and sometimes dominated by wishful thinking. At the least, we should undertake the effort to assess the underlying assertions?including of course those that we advance here?about the degree to which common alleles with usefully important effects exist for chronic disease; such asssessment should use focused and systematic approaches that take the biological realities seriously.
We know these views run against momentum and heavily vested interests. Those interests are promoted by reference to their successes (sometimes overstated, and often argued ?rstor onlythrough the business or journalistic press rather than peer-reviewed articles4). We expect criticism from such quarters, but readers should be aware of the less-publicized failures and inef?ciencies. What per cent of complex disease gene-mapping projects whose grant proposals promised 80% power have actually been successful in identifying the disease genes assumed to be in the data? Application of most current mapping strategies could arguably be more effective if focused on traits that clearly have a genetic basis, such as frank paediatric disorders (which represent a human version of mouse knockouts, where one can assess the effects of the complete absence of a gene product), or on candidate genes, where the genetic or population payoff might be higher, or subsets of complex disease, in samples from appropriately chosen populations where LD may be high and aetiology more marked and homogeneous (or perhaps most importantly through reductions in environmental heterogeneity). Even if such traits are individually rare, they are numerous75, and we know genetic methods work to unravel their aetiologies.
Resistance to genetic reductionism is not new76, and we know that, by expressing these views (some might describe them as heresies), we risk being seen as stereotypic nay-sayers. However, ours is not an argument against genetics, but for a revised genetics that interfaces more intimately with biology. Biological traits have evolved by noise-tolerant evolutionary mechanisms, and a trait that doesnt manifest until long after the reproductive lifespan of most humans throughout history is unlikely to be genetic in the traditional, deterministic sense of the term. Most genetic studies that focus on humans are designed, in effect, to mimic Mendels choice of experimental system, with only two or three frequent states with strongly different effects. That certainly enables us to characterize some of the high-penetrance tail of distribution of the allelic effects, but as noted above these may usually be rather rare. But in?ated claims based on this approach can divert attention from the critical issue of how to deal with complexity on its own terms, and fuel false hopes for simple answers to complex questions. The problems faced in treating complex diseases as if they were Mendels peas show, without invoking the term in its faddish sense, that complexity is a subject that needs its own operating framework, a new twenty-?rst rather than nineteenth?or even twentieth?century genetics.
He even addressed Daniel Macarthur’s potential criticism thirteen years back (shown in bold above).!!
Dr. Weiss continues to hold the same views as you can see from his recent article (April 2013) -
In keeping with the DNA metaphor, the idea of the genomic approach is to assume that genes simply must be causing a trait of interest, and to look across the entire genome to find variants that are more common in individuals with the trait than in those without it. The hope was that we would soon eliminate the debilitating or fatal diseases to which most of us now fall victim, once we had exhaustive knowledge of genome-wide variation.
Genomic studies searching for causal genes have grown ever larger and more expensive, but commensurately important results have yet to roll in. Most of the estimated overall genetic influence on the traits or diseases of interest is still unidentified. What were finding instead is polygenic causation, that is, that many different parts of the genome contribute mainly trivial individual effects.
Each genetic variant is a very weak coin flip with unstable probabilities, and everyone is flipping a different set of coins
A typical well-studied example is Crohns disease, an inflammatory bowel disease that runs in families, and thus would seem to have a major genetic component. However, the most recent study, by Heather Elding and colleagues at University College London, published in The American Journal of Human Genetics, estimates that the number of genes associated with the disease is around 200, most with very small effects, which explains only a small amount of the genetic background of this disease. To liken this again to coin- flipping, variants at each causal gene affect risk in some probabilistic way, usually very small far from 50-50 and with no guarantee whatever that the same variant provides the same risk in different people who carry it, or in different populations, or in men or women, or at different ages. Its as though each coin keeps changing its probability of coming up heads. Thus, the predictive power of this type of personalised genomic medicine is generally very weak, like trying to predict the outcome of hundreds of individual- specific coin-flips. Thats why, with some fortunate exceptions, the clinical or therapeutic value of all these genetic studies has so far been slight.
Its a similar story for normal traits as it is for disease. Height is an easily measured trait that clearly runs in families, and many studies have been done looking for genes for this trait. More than 400 contributing genetic regions, from an estimated 700 or so, have been found but, again, none with very large effects. In fact, to date, only 10 per cent or so of the variation in height has been explained, as a study from Exeter University published in Nature in October 2010 demonstrated. Many more genes will be found to contribute, but environmental factors such as diet or illness will as well.
How does he know so much? For one thing, he happened to write a book titled ‘Genetic Variation and Human Disease: Principles and Evolutionary Approaches’ in 1993. That is 20 years ago !!

Moreover, here is a subset of his pre-1990 papers to show how long he has been thinking about the topics we are discussing today.
Identifying groups at high risk of colorectal cancer - 1983
The genetic structure of a tribal population, the Yanomama Indians. XII. Biodemographic studies.
Genetics and epidemiology of gallbladder disease in New World native peoples.
Epidemiology of breast cancer in a Mexican-American population. - 1985
We do not know, whether he passed Daniel MacArthur’s test for those, who are allowed to criticize GWAS, but here is what Dr. Weiss describes GWAS in his latest blog commentary -
Ranajit Chakraborty and I wrote many papers in the ’80s about the way somatic mutation might explain why cancer is not usually present at birth and to account for the age of onset patterns of cancer, and in my 1993 book Genetic Variation and Human Disease (20 years ago!) and elsewhere I provided a speculative account of how this could apply to age-related diseases (most diseases) more widely.
There were and are many other examples and instances of the importance of somatic mutation, in humans and other animals (and plants). But the bemused human genetics establishment, anchored in early 20th century concepts of simple inheritance, established its juggernaut of GWAS and the idea of relating ‘the’ genome of a person to his/her fate, paying conveniently little attention to somatic mutation.
For context, please check the following two commentaries published in Science:
http://www.sciencemag.org/content/341/6141/1237758.abstract
http://www.sciencemag.org/content/341/6144/358.summary