
Please Try New and Polished HaploMerger2 for Assembly of Heterozygous Genomes
Shengfeng Huang, the author of HaploMerger, informed us about the release of HaploMerger2 built with new and improved algorithm. Those working on assembly of heterozygous genomes should give it a try.
Here is the new flowchart of HaploMerger2 -
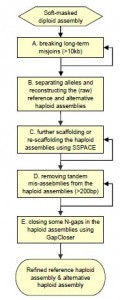
Shengfeng wrote -
Now it comes with a new pipeline which can:
0. be more efficient, accurate and easier to use;
1. reconstruct the allelic relationships within a diploid assembly;
2. detect and correct mis-joins in a diploid assembly;
3. reconstruct two haploid assemblies for the diploid assembly: the reference and the alternative haploid assemblies;
4. further scaffold a haploid assembly (by using third-party software) to restore and achieve better continuity;
5. detect and remove tandem mis-assemblies from haploid (or so diploid) assemblies;
6. finish N-gaps in haploid assemblies (by using third-party software) to achieve better contiguity;
7. Many new features, improved quality and bug fixing.
Download site: http://mosas.sysu.edu.cn/genome/download_softwares.php
-————————————————-
Our previous posts on diploid assembly -
HaploMerger: A Software Tool for Assembling Highly Polymorphic Genomes